1. Introduction
Single-cell sequencing (SCS) seems to be an evolutionary advancement in biomedical sciences, indicating a new understanding of the cellular mechanisms promoting health and disease. This technology expands its analytical reach beyond mere transcriptomics, enabling the research of genetic, epigenetic and proteomic landscapes at the single-cell level. By providing a complex tool for assessing the cellular heterogeneity and dynamics, single-cell sequencing paves the way for advancements in precision medicine. Its capacity to reveal the complexity of individual cells offers an intense departure from traditional bulk sequencing methods, which aggregate genetic signals across cell populations. This advancement in analytical precision plays a crucial role in revealing the complex mechanisms underlying diseases, paving the way for therapeutic interventions that offer greater personalization and improved effectiveness.
Through capturing the genetic and epigenetic variations of individual cells, SCS uncovers the large diversity and complexity within the tissues. This process reveals the distinct cellular dynamics present in various health and disease states, providing a better perspective of the cellular contributions to disease mechanisms and responses to treatments. These insights provided by SCS is essential for examining the complex pathways involved in disease progression and therapeutic reactions, facilitating a progressive change from broad, generalized treatments to more tailored approaches. These approaches, informed by a detailed understanding of an individual's cellular landscape, aim to improve the precision efficacy of medical interventions.
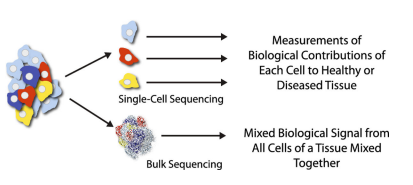
Figure 1. Single cell genome sequencing has higher resolution than bulk sequencing. Single cell genomics enables measurements of biological contributions of each cell to healthy or diseased tissues, whereas bulk sequencing only reflects an average signal of mixed cells.
2. From Bulk to Single-Cell Sequencing: Unlocking Precision Medicine
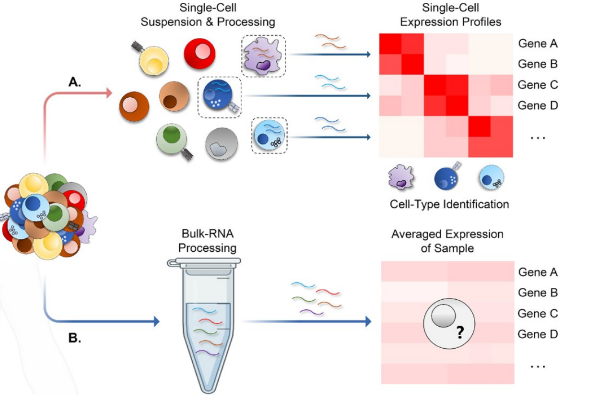
Figure 2. (A) In a single cell data set, the expression data are represented at the individual cell level, providing information on how different cell types impact overall expression. (B) In contrast, in any bulk RNA-Seq, the output data are averaged expression data compared across samples or other types of computational analyses applicable to bulk data.
2.1. Bulk Cell Sequencing
Bulk cell sequencing is a genomic technique where DNA, RNA, or other molecular material is extracted and sequenced from a mixture of many cells at once, rather than from individual cells. This approach provides an averaged molecular profile of the entire cell population in a sample such as a tumor or tissue allowing researchers to analyze common genetic mutations, gene expression patterns, or epigenetic modifications present in the bulk tissue. However, because it combines signals from all cells, bulk sequencing masks the differences between individual cells, making it difficult to detect rare cell types or subpopulations and obscuring cellular heterogeneity within the sample.
2.2. Single Cell Sequencing
Single-cell sequencing is an advanced technology that allows researchers to analyze the genetic, transcriptomic, or epigenetic information of individual cells, rather than averaging signals from a large group of cells as in bulk sequencing. By examining each cell separately, it preserves the unique molecular profile of every single cell in a sample. This high-resolution approach enables the identification of rare cell types, subpopulations, and diverse functional states within complex tissues like tumors. It provides deep insights into cellular heterogeneity, cell-to-cell interactions, and dynamic processes such as tumor evolution and drug resistance.
3. The Problem: Limitations of Bulk Sequencing in Precision Medicine
3.1. Loss of Cellular Heterogeneity
Bulk sequencing aggregates signals from millions of cells, masks rare but clinically significant populations like cancer stem cells or drug resistant cells. Also, struggle to detect low-frequency variants, especially in rare cells (e.g., 1% or less of the tumor), because these signals are mixed in the aggregated data. Bulk RNA-seq cannot differentiate whether gene expression is immune-related or originates from cancer cells or surrounding normal cells. Tumors behave like evolving ecosystems; bulk data can't resolve which mutations co-occur in the same cell or how they expand under treatment pressure, which further limits the insights into clonal evolution and adaptive resistance.
3.2. Inability to Resolve Complex Microenvironments
The tumor microenvironment (TME) is composed of a complex mixture of stromal, immune, and endothelial cells; each of it contributes to disease progression and therapeutic response. However, bulk sequencing techniques hides the complexity by averaging molecular signals across all cell types in a sample. As a result, it becomes difficult to determine whether specific gene expression patterns such as elevated immune signaling are driven by tumor cells themselves or by surrounding immune cells. Moreover, bulk methods are unable to capture the complex intercellular interactions within the TME, which are essential for understanding key processes like immune evasion, angiogenesis, and metastasis. This limitation inhibits a comprehensive and detailed understanding of tumor biology and its microenvironment.
3.3. Hidden Clonal Evolution and Treatment Resistance
Bulk sequencing captures only an averaged snapshot of the genomic landscape, offering neither temporal nor spatial resolution. This limitation makes it insufficient for revealing the dynamic evolutionary processes occurring within tumors, such as an expansion of distinct subclonal populations. Subclonal mapping from bulk data can lead to significant inaccuracies, particularly when smaller clones are represented by low-frequency variants that are easily dominated or misclassified. As tumors evolve under therapeutic pressure, resistant subclones can develop and drive treatment failure. However, these critical clones often go unnoticed in bulk sequencing due to their unusual emergence. Clinical studies, including those on chronic myeloid leukemia, have shown that such undetected cells can eventually dominate and lead to disease reversion highlighting a major gap in bulk sequencing's ability to inform timely and effective treatment strategies.
4. The Solution: Single-Cell Sequencing (SCS)
4.1. Resolving Cellular Heterogeneity
In contrast to bulk sequencing, which blends molecular signals from millions of cells into a single average profile, single-cell sequencing analyzes each cell individually, preserving its unique molecular characteristics. This enables the identification of rare but clinically significant subpopulations, such as cancer stem cells, drug resistant clones, or dormant cells that may later contribute to disease relapse. By capturing cell type specific mutations, epigenetic modifications, and gene expression patterns, single-cell sequencing provides a far more precise and extensive view of tissue heterogeneity. It also allows researchers to map the transcriptional diversity within tumors, uncovering distinct functional states such as proliferative, invasive, or immune evasive phenotypes that are essential for developing personalized treatment strategies.
4.2. Decoding the Tumor Microenvironment (TME)
Single-cell technologies offer high-resolution insights into the tumor microenvironment (TME) by comparing between cancer cells and surrounding stromal, immune, and endothelial cells. This detailed resolution allows researchers to evaluate complex cell to cell interactions, including immune-tumor signaling, cytokine activity, and stromal support dynamics. Additionally, single cell approaches enable comprehensive immune profiling for example, identifying specific T cell subtypes or macrophage position states which is crucial for designing effective immunotherapy strategies. When integrated with transcriptomics, these technologies not only reveal the types of cells present but also map their precise locations within the tissue, providing a more complete understanding of the TME's structure and function.
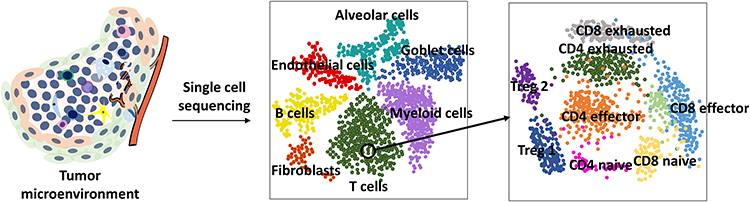
4.3. Tracking Clonal Evolution and Drug Resistance
Single cell DNA and RNA sequencing enables the tracking of tumor development by capturing the emergence and growth of treatment resistant cells. By comparing the mutational profiles of individual cells, these techniques allow the reconstruction of detailed phylogenetic trees that illustrate the cancer's evolutionary history. They also observe clonal fluctuation throughout the course of therapy, revealing how particular subclones survive treatment and expand. This level of resolution facilitates the early detection of resistance mechanisms, permitting clinicians to adjust treatment strategies proactively before relapse occurs.
5. How Single-Cell Sequencing Overcomes Bulk Sequencing Limitation
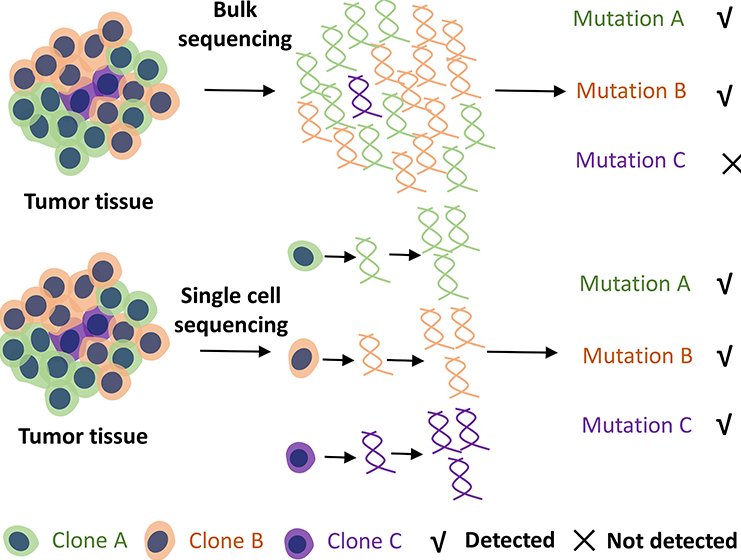
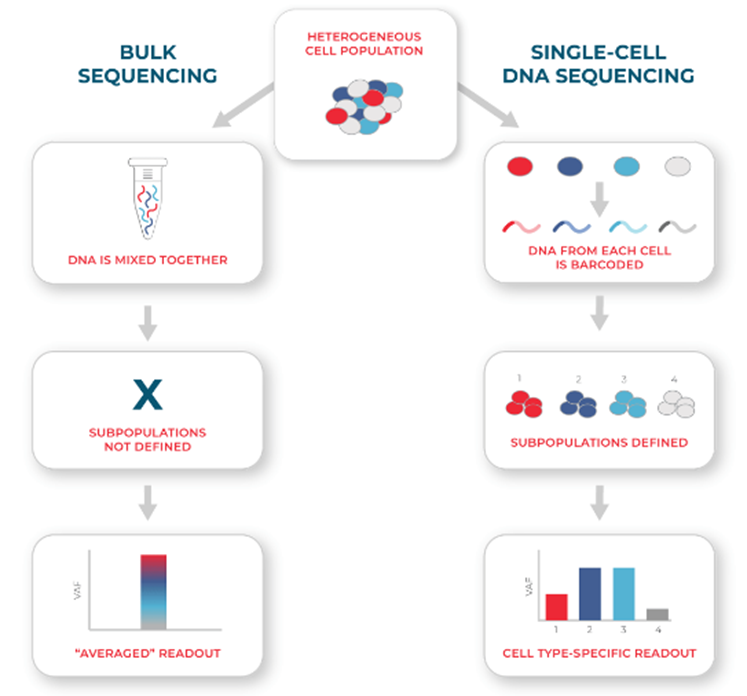
Figure 3. With the single-cell resolution of the technique, single-cell sequencing can assess heterogeneity better and be more sensitive for detecting rare mutations than conventional bulk sequencing.
| Challenge | Bulk Sequencing Limitation | Single Cell Sequencing Solution |
|---|---|---|
| Rare Subpopulation Detection | Masked by averaging over many cells | Resolves even very rare cell types and states |
| Tumor Microenvironment Analysis | Mixed cell populations indistinguishable | Identifies and characterizes distinct cell types |
| Clonal Evolution Tracking | Unable to detect subclonal dynamics | Traces lineage evolution and mutation acquisition over time |
| Cellular Heterogeneity | Remains unknown by bulk measurements | Provides a detailed map of cellular diversity and plasticity |
6. Driving Precision Medicine with Single-Cell Technologies
Single-cell technologies are transforming precision medicine by revealing the molecular details of individual cells within the complex tissues. Unlike bulk sequencing, they detect cellular heterogeneity, track disease progression at the clonal level, and identify rare cell types that influence treatment response. This enables more accurate diagnostics and personalized therapies tailored to each patient's unique cellular environment.
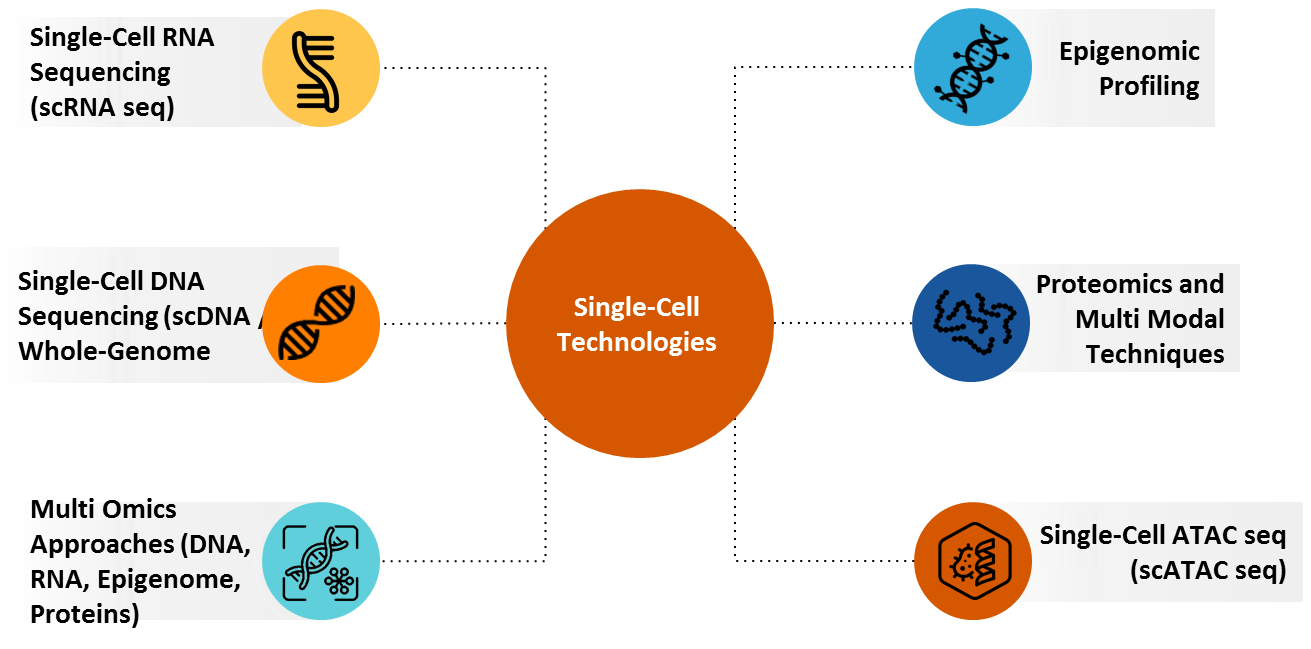
6.1. Single-Cell RNA Sequencing (scRNA seq)
Single-cell RNA sequencing (scRNA-seq) is an important technique that analyzes gene expression at the individual cell level, revealing cellular diversity hidden in bulk analysis. It allows precise identification of rare cell types, disease-related cell states, and dynamic biological processes. In precision medicine, scRNA-seq helps tailor treatments by revealing patient-specific cellular profiles, especially in cancer, immune, and neurological disorders. The technology advances personalized therapies by identifying novel biomarkers and therapeutic targets. Its integration into clinical research is transforming diagnostics, drug development, and individualized care strategies.
6.2. Single-Cell DNA Sequencing (scDNA seq) / Whole-Genome
Single-cell DNA sequencing (scDNA-seq) allows the analysis of genomic variations at the single-cell level, revealing genetic heterogeneity within tissues. It detects mutations, copy number variations (CNVs), and structural changes that are often missed in bulk sequencing. In precision medicine, scDNA-seq helps in identifying rare cancer subclones, track tumor evolution, and monitor treatment resistance. It is crucial for understanding the genetic basis of complex diseases and tailoring targeted therapies. As technology advances, scDNA-seq is increasingly used in personalized diagnostics and clinical decision-making.
6.3. Multi Omics Approaches (DNA, RNA, Epigenome, Proteins)
Multi-omics approaches in single-cell sequencing integrate data from DNA, RNA, epigenome, and proteins to provide a complete molecular profile of individual cells. This enables a deeper understanding of how genetic mutations, gene expression, epigenetic regulation, and protein activity interact within each cell. In precision medicine, it helps identify disease-driving cell subpopulations, predict treatment responses, and uncover personalized therapeutic targets.
6.4. Epigenomic Profiling
Single-cell epigenomic profiling analyzes DNA methylation, chromatin accessibility, and histone modifications at the individual cell level, revealing how gene expression is regulated beyond DNA sequence. Further, it reveals cell-specific regulatory mechanisms and epigenetic heterogeneity in health and disease. In precision medicine, this helps identify disease-associated epigenetic changes, especially in cancer, neurodegeneration, and immune disorders. Techniques like scATAC-seq and scBS-seq enable high-resolution mapping of the epigenome. This approach supports personalized therapy by linking epigenetic states to treatment response and disease progression.
6.5. Proteomics and Multi Modal Techniques
Single-cell proteomics and multi-modal techniques analyze protein expression with other molecular layers (like RNA or chromatin) within the individual cells. These approaches provide direct insight into cell signaling, functional states, and post-translational modifications that RNA data alone cannot reveal. Techniques like CITE-seq and REAP-seq combine transcriptomics and surface protein profiling, enhancing cellular phenotyping in diseases. In precision medicine, they help identify functional biomarkers, drug targets, and treatment-responsive cell types. This integrative approach enables more accurate disease classification and tailored therapeutic strategies.
6.6. Single-Cell ATAC seq (scATAC seq)
Single-cell ATAC-seq (scATAC-seq) profiles chromatin accessibility at the individual cell level, identifying open regions of the genome where gene regulation occurs. This technique reveals cell-specific regulatory elements and transcription factor activity that drive gene expression patterns. In precision medicine, scATAC-seq helps uncover epigenetic heterogeneity in diseases like cancer, aiding in the identification of disease-driving cell populations. It supports the discovery of novel biomarkers and therapeutic targets by linking chromatin states to cellular function. Overall, scATAC-seq enhances understanding of gene regulation dynamics for personalized diagnosis and treatment.
| Technique | What it Measures |
|---|---|
| Single-Cell RNA Sequencing (scRNA seq) | Gene expression at single-cell resolution |
| Single-Cell DNA Sequencing (scDNA seq) / Whole-Genome | Genomic mutation profiling in individual cells |
| Multi Omics Approaches (DNA, RNA, Epigenome, Proteins) | Integrated genome, transcriptome, epigenome data |
| Epigenomic Profiling | DNA methylation, histone modifications, chromatin structure |
| Proteomics and Multi Modal Techniques | Protein and mRNA quantification in single cells |
| Single-Cell ATAC seq (scATAC seq) | Chromatin accessibility and regulatory element profiling |
7. Key Innovation of Single Cell Sequencing in precision medicine
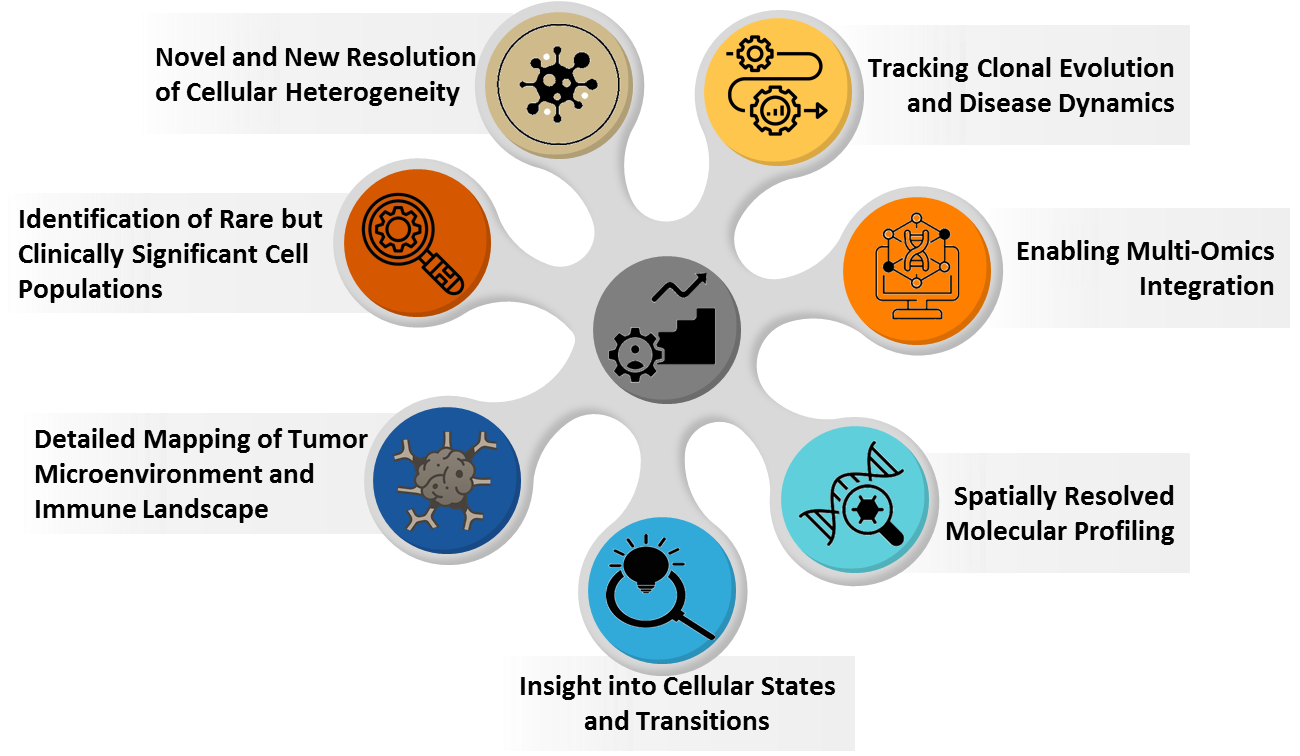
8. Key application of Single Cell Sequencing in precision medicine
1. Oncology: Detection of minimal residual disease and rare resistant clones to guide therapy modifications. Characterization of tumor immune cells to predict immunotherapy responses. Understanding metastatic mechanisms through detailed cell lineage tracing.
2. Immunology and Autoimmune Diseases: Profiling immune cell diversity and activation states at single-cell resolution. Identifying pathogenic cell subsets responsible for autoimmune pathology.
3. Neurology: Mapping neuronal and glial cell heterogeneity in neurodegenerative diseases. Solves disease mechanisms at a cellular level for targeted interventions.
4. Infectious Diseases: Profiling infected versus uninfected cells in heterogeneous tissues. Understanding host-pathogen interactions and immune evasion strategies.
9. Future Directions
As single-cell sequencing technologies continue to evolve, they are expected to become more affordable, scalable, and integrated into routine clinical practice. Future advancements will likely include real-time single-cell diagnostics, enhanced multi-omics platforms for faster and more accurate interpretation of complex cellular data. These innovations will enhance the understanding of disease mechanisms, enable earlier detection of treatment resistance, and guide highly personalized therapeutic interventions. In the coming years, SCS will play a primary role in next-generation precision medicine, transforming how we diagnose, treat, and monitor a wide range of diseases. The integration of single-cell data into electronic health records may also revolutionize patient care and clinical conclusion.
10. Conclusion
In conclusion, single-cell sequencing (SCS) represents a revolutionary breakthrough in biomedical research and precision medicine, offering new and novel insights into cellular heterogeneity, tissue complexity, and disease progression. By exceeding the limitations of traditional bulk sequencing, SCS allows for the detailed profiling of individual cells, enabling the detection of rare subpopulations, tracking of clonal evolution, and decoding of the tumor microenvironment. Techniques such as scRNA-seq, scDNA-seq, scATAC-seq, and multi-omics approaches provide a complete view of gene expression, genomic mutations, epigenetic regulation, and protein activity at the single-cell level. These innovations enables researchers and clinicians to develop more personalized and effective diagnostic, therapeutic, and monitoring strategies across a range of diseases, including cancer, autoimmune disorders, neurological conditions, and infections. As single-cell technologies continue to evolve and integrate, they will play an increasingly central role in shaping the future of individualized patient care.




-Innovative-Applications-and-Insights.jpeg)






